
It is only recently, with the advent and proliferation of NGS technology, have we been able to fully take advantage of RNA-seq’s potential. 2Įarly RNA-seq techniques used Sanger sequencing technology, a technique that although innovative at the time was also low-throughput and costly. It can also identify post-transcriptional modifications that occur during mRNA processing such as polyadenylation and 5’ capping. These events would not be picked up by DNA sequencing. It also captures information about alternative splicing events (Figure 1), which produce different transcripts from one single gene sequence. For example, the transcriptome can highlight all the tissues in which a gene of unknown function is turned on, which might indicate what its role is. This can give researchers vital information about the function of genes. Some of the most popular techniques that use RNA-seq are transcriptional profiling, single nucleotide polymorphism (SNP) identification, 3 RNA editing and differential gene expression analysis. 2 This allows scientists to understand the biology of a cell more deeply and assess changes that may indicate disease. RNA-seq can tell us which genes are turned on in a cell, what their level of transcription is, and at what times they are activated or shut off. 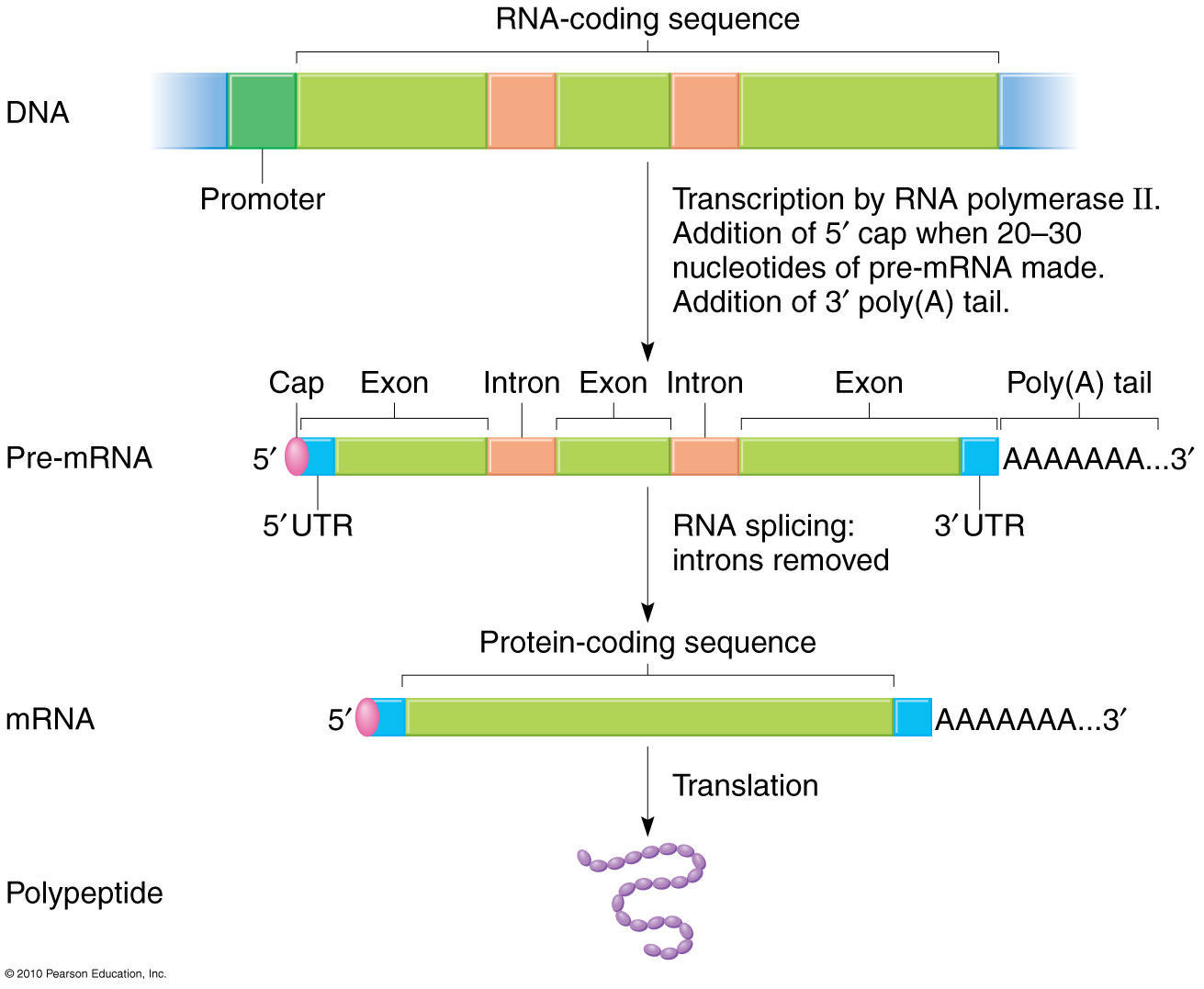
Understanding the transcriptome is key if we are to connect the information in our genome with its functional protein expression. RNA-seq lets us investigate and discover the transcriptome, the total cellular content of RNAs including mRNA, rRNA and tRNA.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
